When training models and when applying Expressions (e.g., Classification using expression) the spectrum can benefit from applying a pretreatment. Here we will expand on the use-case for each available pretreatment.
Savitzky-Golay
Savitzky-Golay is based on a moving window that fit a polynomial curve of fixed degree to the spectral data. This implies that the signal for each observation is smoothed according to a window size that the user can specify. The user must give the polynomial order of the equation that is to be used for the smoothing. A higher polynomial order will ensure a better fit. The user also has to give the wanted derivative order. Please note that the derivative order cannot be higher than the polynomial order. Finally, the user has to give the number of left and right points, which is equivalent to the window size used for the smoothing.

Read more at: https://en.wikipedia.org/wiki/Savitzky%E2%80%93Golay_filter
Continuum Removal
Continuum Removal involves identifying the spectral continuum - a baseline connecting the highest points in a spectral curve - and normalizing the data by dividing it with this continuum. This process highlights specific absorption features in the spectral data, making it easier to compare and identify different materials or conditions.
Left Offset and Right Offset define the wavelength range around each absorption feature that you are interested in analyzing. The left offset specifies the starting point (in wavelengths) to the left of the absorption feature, and the right offset specifies the ending point to the right. Adjusting these offsets allows you to focus on specific parts of the spectral data, ensuring that the continuum is accurately identified and the absorption features are properly isolated for your analysis. This customization is crucial for accurate material identification and comparative studies in various applications like environmental monitoring or geological mapping.

Derivative
Calculates the first-derivative or second-derivative for all observations. This is useful as it removes constant background signals.
Tip
This may work well if your are using quantification with small variations
SNV (Standard Normal Variate correction)
The so-called Standard Normal Variate (SNV) method performs a normalization of the spectra that consists in subtracting each spectrum by its own mean and dividing it by its own standard deviation. After SNV, each spectrum will have a mean of 0 and a standard deviation of 1.
Tip
This can be used to remove scattering effects caused by physical variations such as height differences and and surface imperfections
Logarithm
Adds the logarithms of the selected variables. The user can enter a constant in the settings panel in order to prevent that logarithms are performed for values of zero or less.
Center
By default, Breeze uses variable mean centering when preparing a DataSet prior to creating a multivariate PCA or PLS model. In variable mean centering the mean value of a variable (i.e., the mean value of all observations) is subtracted from each individual value. Visually, this can be seen as the center of the multidimensional variable space is shifted towards the origin of coordinates. Variable mean centering implies that the subsequent multivariate modeling is more straightforward, as the variable means, if not removed, would have added one component to the model. This simplifies both the model complexity and the interpretation of data.
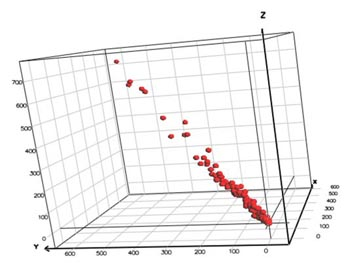
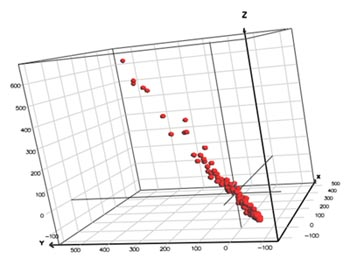
UV (Unit Variance) scaling
UV scaling means that all variables are divided by their standard deviations. This implies that all variables have equal variation of one unit. For certain kinds of data, e.g. when variables have different units, this can improve the modeling as some variables with large variation would otherwise influence the models. For other kinds of data, e.g. when variables consists of spectral measurements, unit variance scaling will make the noise more prominent, thus making the modeling more difficult.
UV scaling is almost always used in conjunction with variable mean Pretreatments | Center
Can increase noise in the model
